Biosensing detection system for detecting ctDNA and detection method thereof
A detection system and biosensing technology, applied in biochemical equipment and methods, microbial determination/inspection, etc., can solve the problems of inability to detect clinical low-abundance samples, low sensitivity, poor stability, etc., and achieve excellent resistance to nucleic acids. Enzymatic cleavage, improved sensitivity and improved stability
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
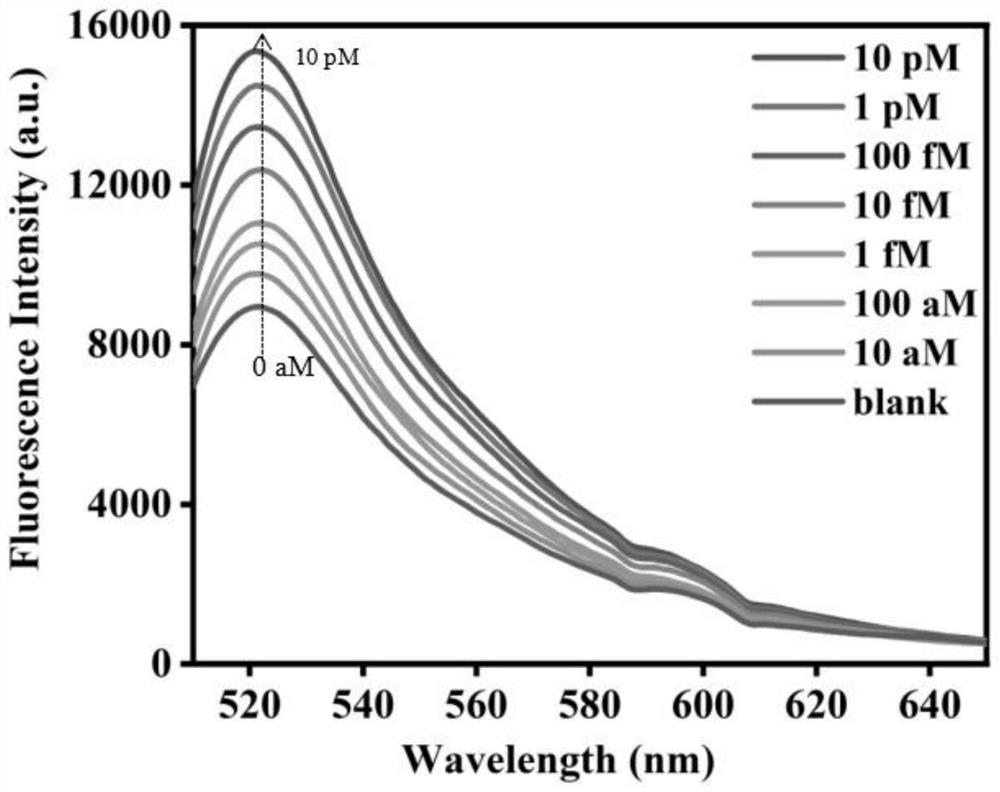
[0026] Example 1 - Biosensor Sensitivity Analysis
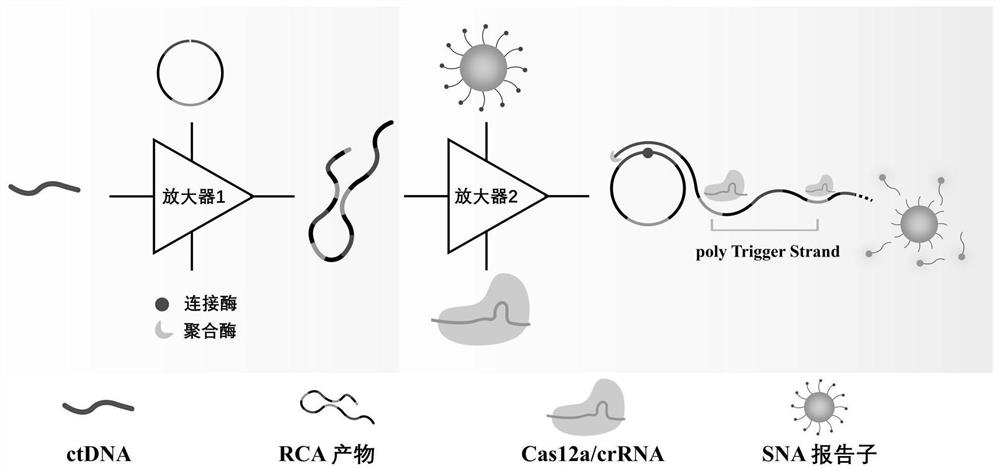
[0027] Detection method flow chart of the present invention, as figure 1 As shown, the specific steps are as follows:
specific Embodiment approach
[0028] This embodiment provides a specific implementation of a biosensor based on ctDNA highly sensitive and specific detection, specifically the following steps:
[0029] S1. Preparation of 13nm gold nanoparticles: first soak the round bottom flask with aqua regia, and then thoroughly clean it with ultrapure water. Add 100mL of 0.01wt% chloroauric acid solution to a clean round bottom flask, then put the round bottom flask into an oil bath, set the oil bath temperature to 130°C, and the rotation speed to 800rpm. After the solution boils, continue to boil for 10 minutes to remove the dissolved oxygen in the solution. Increase the rotation speed to 1200rpm, then quickly add 1mL of 3wt% sodium citrate to the solution, the solution changes from light yellow to dark gray, then purple red, and finally wine red, and continue to boil after the color remains unchanged 30min. After the solution was naturally cooled to room temperature, it was stored at 4° C. in the dark, and the average s
Embodiment 2
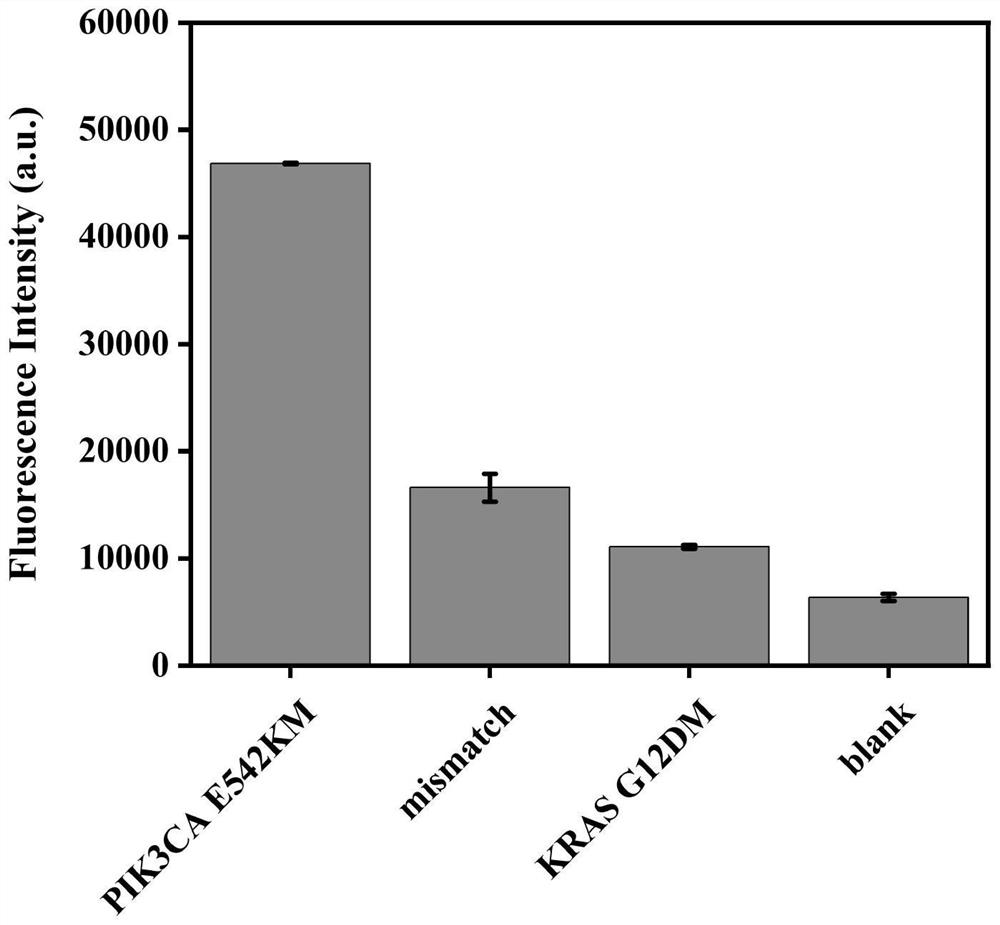
[0037] The difference between the steps of this example and Example 1 lies in the rolling circle amplification reaction stage. In this example, 2 μL of 100 nM 5’ phosphorylated linear padlock probe, 2 μL of different types of ctDNA (PIK3CAE542KM, mismatch-PIK3CAE542KM, KRAS G12DM) and 0.3 μL of T4 ligase (1000 U / μL) were mixed in 1×T4 Incubate at 20°C for 20 minutes in ligase reaction buffer, and then heat-treat at 65°C for 5 minutes to inactivate T4 ligase, and finally obtain a circular DNA template. In the amplification reaction, add 8 μL dNTP (2.5 mM), 0.8 μL BSA (20 mg / mL), 0.4 μL phi29 enzyme (10 U / μL) and 4 uL phi29 buffer to the above reaction solution, incubate at 30°C for 30 minutes, and heat at 65°C for 10 minutes. Finally, the long ssDNA product, namely the RCA product, was obtained.
[0038] The selective analysis results of the biosensing detection system are as follows: image 3 As shown, the RCA-CRISPR / Cas12a system only has a significant fluorescence res
PUM
| Property | Measurement | Unit |
|---|---|---|
| Average size | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to view more
Login to view more - R&D Engineer
- R&D Manager
- IP Professional
- Industry Leading Data Capabilities
- Powerful AI technology
- Patent DNA Extraction
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic.
© 2024 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap